In clinical in vitro fertilization (IVF), the prevailing method for preimplantation genetic testing for aneuploidy (PGT-A) requires biopsying a few cells from the trophectoderm (TE), the lineage that forms the placenta. The test is invasive, requires specialized skills, and suffers from false positives and negatives (Figure 1) because chromosome numbers different from the inner cell mass (ICM) that develops into the fetus.
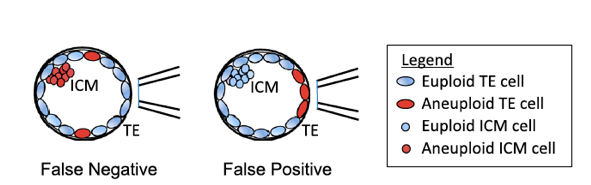
Fig. 1.
False positives and false negatives arise from embryo mosaicism in TE biopsy PGT-A.
Noninvasive PGT-A (niPGT-A), which is based on sequencing DNA released into the culture medium from both TE and ICM, may offer a solution to these problems but has previously had limited efficacy (Xu et al., 2016).
In this study, fifty-two frozen donated blastocysts with TE biopsy results were thawed; each of their spent culture medium was collected after 24-h culture and analyzed by next-generation sequencing (NGS). niPGT-A and TE-biopsy PGT-A results were compared with the sequencing results of the corresponding embryos, which were taken as true results for aneuploidy reporting (Figure 2).
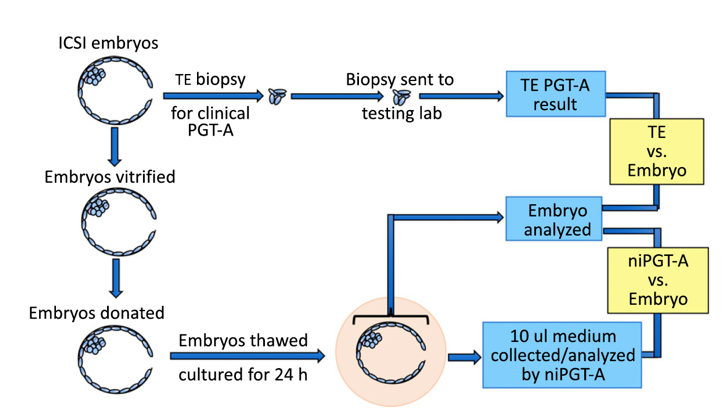
Fig. 2.
Workflow of sample processing for PGT-A analysis of TE biopsy, embryo, and spent culture media.
We note that the fact that the Y chromosomes have a copy number (CN) of one in our niPGT-A data (Figure 3A, 3B) indicates the lack of maternal contamination from the cumulus cells in the culture media, which is essential for the ploidy detection. Among the 52 niPGT-A CN profiles, 4 showed numerous chromosomes with multiple gains and losses, and have high coefficients of variation (CVs), they were categorized as “noisy trace” (Figure 3C). An example of a mosaic embryo in the embryo CN profile and in the niPGT-A CN profile were shown in Figure 3D.
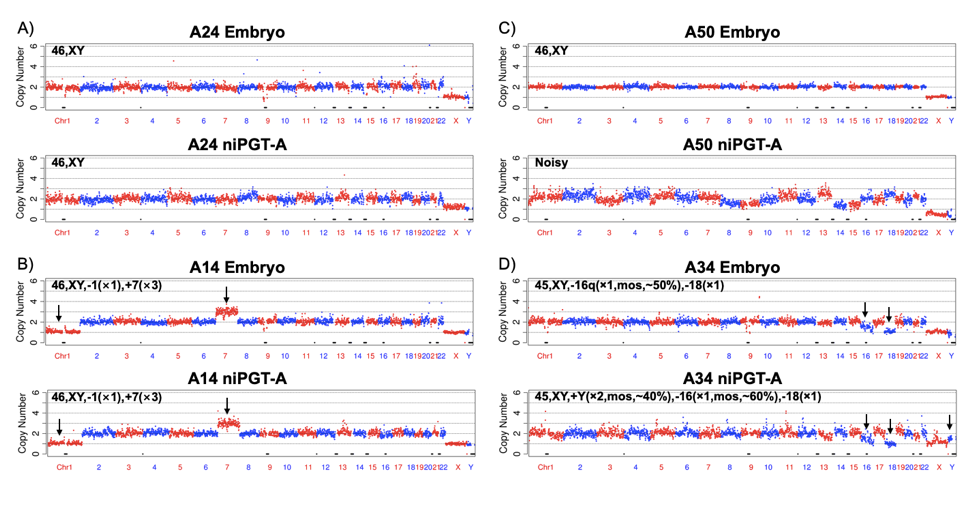
Fig. 3.
The CN plots of embryo vs. spent culture medium at a sequencing depth of 0.02x. The chromosomes are shown in alternating red and blue colors.
To distinguish true embryo aneuploidy from the noise associated with WGA and biological clearing of aneuploid cells, a threshold for mosaicism had to be carefully selected to minimize the false positives and false negatives. The FPR and FNR detected by niPGT-A were used as a function of the mosaicism threshold (Figure 4). We chose 60% as the threshold for distinguishing aneuploid from euploid embryos.
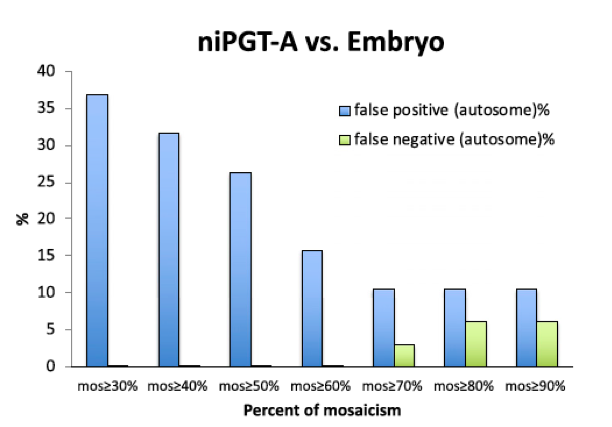
Fig. 4.
FPR and FNR as a function of the percent mosaicism in niPGT-A profiles. Sixty percent of mosaicism was set as the threshold for identifying aneuploidy.
By applying an appropriate threshold for mosaicism, both the positive predictive value (PPV) and specificity for niPGT-A were much higher than TE-biopsy PGT-A. With removal of all corona-cumulus cells, the false-negative rate (FNR) for niPGT-A was found to be zero. Furthermore, the concordance rates for both embryo ploidy and chromosome copy numbers were higher for niPGT-A than TE-biopsy PGT-A. These results suggest that niPGT-A is less prone to errors caused by potential mosaicism in the blastocyst, and is a more reliable test than the current testing method, trophectoderm biopsy.
References:
Huang, Lei; Bogale, Berhan; Tang, Yaqiong; Lu, Sijia; Xie, X. Sunney; Racowsky, Catherine "Non-invasive Preimplantation Genetic Testing for Aneuploidy (niPGT-A) in Spent Medium may be More Reliable than Trophectoderm Biopsy," Proc Natl Acad Sci USA (2019)
Xu, Juanjuan; Fang, Rui; Chen, Li; Chen, Daozhen; Xiao, Jian-Ping; Yang, Weimin; Wang, Honghua; Song, Xiaoqing; Ma, Ting; Bo, Shiping; Shi, Chong; Ren, Jun; Huang, Lei; Cai, Li-Yi; Yao, Bing; Xie, X Sunney; Lu, Sijia "Noninvasive chromosome screening of human embryos by genome sequencing of embryo culture medium for in vitro fertilization," Proc Natl Acad Sci USA 113(42), 11907-11912. DOI: 10.1073/pnas.1613294113 (2016)
|

